About six months in the past, we confirmed create a customized wrapper to acquire uncertainty estimates from a Keras community. Right this moment we current a much less laborious, as properly faster-running means utilizing tfprobability, the R wrapper to TensorFlow Likelihood. Like most posts on this weblog, this one received’t be quick, so let’s shortly state what you possibly can anticipate in return of studying time.
What to anticipate from this publish
Ranging from what not to anticipate: There received’t be a recipe that tells you the way precisely to set all parameters concerned with a view to report the “proper” uncertainty measures. However then, what are the “proper” uncertainty measures? Except you occur to work with a technique that has no (hyper-)parameters to tweak, there’ll all the time be questions on report uncertainty.
What you can anticipate, although, is an introduction to acquiring uncertainty estimates for Keras networks, in addition to an empirical report of how tweaking (hyper-)parameters could have an effect on the outcomes. As within the aforementioned publish, we carry out our assessments on each a simulated and an actual dataset, the Mixed Cycle Energy Plant Information Set. On the finish, instead of strict guidelines, it’s best to have acquired some instinct that can switch to different real-world datasets.
Did you discover our speaking about Keras networks above? Certainly this publish has a further aim: Up to now, we haven’t actually mentioned but how tfprobability goes along with keras. Now we lastly do (in brief: they work collectively seemlessly).
Lastly, the notions of aleatoric and epistemic uncertainty, which can have stayed a bit summary within the prior publish, ought to get way more concrete right here.
Aleatoric vs. epistemic uncertainty
Reminiscent by some means of the traditional decomposition of generalization error into bias and variance, splitting uncertainty into its epistemic and aleatoric constituents separates an irreducible from a reducible half.
The reducible half pertains to imperfection within the mannequin: In idea, if our mannequin have been good, epistemic uncertainty would vanish. Put in another way, if the coaching information have been limitless – or in the event that they comprised the entire inhabitants – we might simply add capability to the mannequin till we’ve obtained an ideal match.
In distinction, usually there may be variation in our measurements. There could also be one true course of that determines my resting coronary heart charge; nonetheless, precise measurements will fluctuate over time. There’s nothing to be completed about this: That is the aleatoric half that simply stays, to be factored into our expectations.
Now studying this, you is perhaps pondering: “Wouldn’t a mannequin that really have been good seize these pseudo-random fluctuations?”. We’ll depart that phisosophical query be; as a substitute, we’ll attempt to illustrate the usefulness of this distinction by instance, in a sensible means. In a nutshell, viewing a mannequin’s aleatoric uncertainty output ought to warning us to consider acceptable deviations when making our predictions, whereas inspecting epistemic uncertainty ought to assist us re-think the appropriateness of the chosen mannequin.
Now let’s dive in and see how we could accomplish our aim with tfprobability. We begin with the simulated dataset.
Uncertainty estimates on simulated information
Dataset
We re-use the dataset from the Google TensorFlow Likelihood workforce’s weblog publish on the identical topic , with one exception: We lengthen the vary of the impartial variable a bit on the damaging aspect, to higher reveal the completely different strategies’ behaviors.
Right here is the data-generating course of. We additionally get library loading out of the way in which. Just like the previous posts on tfprobability, this one too options lately added performance, so please use the event variations of tensorflow and tfprobability in addition to keras. Name install_tensorflow(model = "nightly") to acquire a present nightly construct of TensorFlow and TensorFlow Likelihood:
# be certain we use the event variations of tensorflow, tfprobability and keras
devtools::install_github("rstudio/tensorflow")
devtools::install_github("rstudio/tfprobability")
devtools::install_github("rstudio/keras")
# and that we use a nightly construct of TensorFlow and TensorFlow Likelihood
tensorflow::install_tensorflow(model = "nightly")
library(tensorflow)
library(tfprobability)
library(keras)
library(dplyr)
library(tidyr)
library(ggplot2)
# be certain this code is appropriate with TensorFlow 2.0
tf$compat$v1$enable_v2_behavior()
# generate the information
x_min <- -40
x_max <- 60
n <- 150
w0 <- 0.125
b0 <- 5
normalize <- operate(x) (x - x_min) / (x_max - x_min)
# coaching information; predictor
x <- x_min + (x_max - x_min) * runif(n) %>% as.matrix()
# coaching information; goal
eps <- rnorm(n) * (3 * (0.25 + (normalize(x)) ^ 2))
y <- (w0 * x * (1 + sin(x)) + b0) + eps
# take a look at information (predictor)
x_test <- seq(x_min, x_max, size.out = n) %>% as.matrix()How does the information look?
ggplot(information.body(x = x, y = y), aes(x, y)) + geom_point()
Determine 1: Simulated information
The duty right here is single-predictor regression, which in precept we will obtain use Keras dense layers.
Let’s see improve this by indicating uncertainty, ranging from the aleatoric kind.
Aleatoric uncertainty
Aleatoric uncertainty, by definition, just isn’t an announcement in regards to the mannequin. So why not have the mannequin be taught the uncertainty inherent within the information?
That is precisely how aleatoric uncertainty is operationalized on this method. As a substitute of a single output per enter – the expected imply of the regression – right here we’ve got two outputs: one for the imply, and one for the usual deviation.
How will we use these? Till shortly, we might have needed to roll our personal logic. Now with tfprobability, we make the community output not tensors, however distributions – put in another way, we make the final layer a distribution layer.
Distribution layers are Keras layers, however contributed by tfprobability. The superior factor is that we will prepare them with simply tensors as targets, as normal: No must compute chances ourselves.
A number of specialised distribution layers exist, corresponding to layer_kl_divergence_add_loss, layer_independent_bernoulli, or layer_mixture_same_family, however probably the most normal is layer_distribution_lambda. layer_distribution_lambda takes as inputs the previous layer and outputs a distribution. So as to have the ability to do that, we have to inform it make use of the previous layer’s activations.
In our case, sooner or later we’ll need to have a dense layer with two models.
... %>% layer_dense(models = 2, activation = "linear") %>%Then layer_distribution_lambda will use the primary unit because the imply of a standard distribution, and the second as its customary deviation.
layer_distribution_lambda(operate(x)
tfd_normal(loc = x[, 1, drop = FALSE],
scale = 1e-3 + tf$math$softplus(x[, 2, drop = FALSE])
)
)Right here is the entire mannequin we use. We insert a further dense layer in entrance, with a relu activation, to provide the mannequin a bit extra freedom and capability. We focus on this, in addition to that scale = ... foo, as quickly as we’ve completed our walkthrough of mannequin coaching.
mannequin <- keras_model_sequential() %>%
layer_dense(models = 8, activation = "relu") %>%
layer_dense(models = 2, activation = "linear") %>%
layer_distribution_lambda(operate(x)
tfd_normal(loc = x[, 1, drop = FALSE],
# ignore on first learn, we'll come again to this
# scale = 1e-3 + 0.05 * tf$math$softplus(x[, 2, drop = FALSE])
scale = 1e-3 + tf$math$softplus(x[, 2, drop = FALSE])
)
)For a mannequin that outputs a distribution, the loss is the damaging log chance given the goal information.
negloglik <- operate(y, mannequin) - (mannequin %>% tfd_log_prob(y))We will now compile and match the mannequin.
We now name the mannequin on the take a look at information to acquire the predictions. The predictions now truly are distributions, and we’ve got 150 of them, one for every datapoint:
yhat <- mannequin(tf$fixed(x_test))tfp.distributions.Regular("sequential/distribution_lambda/Regular/",
batch_shape=[150, 1], event_shape=[], dtype=float32)To acquire the means and customary deviations – the latter being that measure of aleatoric uncertainty we’re all in favour of – we simply name tfd_mean and tfd_stddev on these distributions.
That can give us the expected imply, in addition to the expected variance, per datapoint.
Let’s visualize this. Listed here are the precise take a look at information factors, the expected means, in addition to confidence bands indicating the imply estimate plus/minus two customary deviations.
ggplot(information.body(
x = x,
y = y,
imply = as.numeric(imply),
sd = as.numeric(sd)
),
aes(x, y)) +
geom_point() +
geom_line(aes(x = x_test, y = imply), shade = "violet", dimension = 1.5) +
geom_ribbon(aes(
x = x_test,
ymin = imply - 2 * sd,
ymax = imply + 2 * sd
),
alpha = 0.2,
fill = "gray")
Determine 2: Aleatoric uncertainty on simulated information, utilizing relu activation within the first dense layer.
This appears to be like fairly cheap. What if we had used linear activation within the first layer? Which means, what if the mannequin had appeared like this:
This time, the mannequin doesn’t seize the “kind” of the information that properly, as we’ve disallowed any nonlinearities.

Determine 3: Aleatoric uncertainty on simulated information, utilizing linear activation within the first dense layer.
Utilizing linear activations solely, we additionally must do extra experimenting with the scale = ... line to get the consequence look “proper”. With relu, however, outcomes are fairly sturdy to adjustments in how scale is computed. Which activation can we select? If our aim is to adequately mannequin variation within the information, we will simply select relu – and depart assessing uncertainty within the mannequin to a distinct method (the epistemic uncertainty that’s up subsequent).
Total, it looks like aleatoric uncertainty is the easy half. We wish the community to be taught the variation inherent within the information, which it does. What can we achieve? As a substitute of acquiring simply level estimates, which on this instance would possibly prove fairly unhealthy within the two fan-like areas of the information on the left and proper sides, we be taught in regards to the unfold as properly. We’ll thus be appropriately cautious relying on what enter vary we’re making predictions for.
Epistemic uncertainty
Now our focus is on the mannequin. Given a speficic mannequin (e.g., one from the linear household), what sort of information does it say conforms to its expectations?
To reply this query, we make use of a variational-dense layer.
That is once more a Keras layer offered by tfprobability. Internally, it really works by minimizing the proof decrease sure (ELBO), thus striving to search out an approximative posterior that does two issues:
- match the precise information properly (put in another way: obtain excessive log chance), and
- keep near a prior (as measured by KL divergence).
As customers, we truly specify the type of the posterior in addition to that of the prior. Right here is how a previous might look.
prior_trainable <-
operate(kernel_size,
bias_size = 0,
dtype = NULL) {
n <- kernel_size + bias_size
keras_model_sequential() %>%
# we'll touch upon this quickly
# layer_variable(n, dtype = dtype, trainable = FALSE) %>%
layer_variable(n, dtype = dtype, trainable = TRUE) %>%
layer_distribution_lambda(operate(t) {
tfd_independent(tfd_normal(loc = t, scale = 1),
reinterpreted_batch_ndims = 1)
})
}This prior is itself a Keras mannequin, containing a layer that wraps a variable and a layer_distribution_lambda, that kind of distribution-yielding layer we’ve simply encountered above. The variable layer may very well be fastened (non-trainable) or non-trainable, similar to a real prior or a previous learnt from the information in an empirical Bayes-like means. The distribution layer outputs a standard distribution since we’re in a regression setting.
The posterior too is a Keras mannequin – positively trainable this time. It too outputs a standard distribution:
posterior_mean_field <-
operate(kernel_size,
bias_size = 0,
dtype = NULL) {
n <- kernel_size + bias_size
c <- log(expm1(1))
keras_model_sequential(record(
layer_variable(form = 2 * n, dtype = dtype),
layer_distribution_lambda(
make_distribution_fn = operate(t) {
tfd_independent(tfd_normal(
loc = t[1:n],
scale = 1e-5 + tf$nn$softplus(c + t[(n + 1):(2 * n)])
), reinterpreted_batch_ndims = 1)
}
)
))
}Now that we’ve outlined each, we will arrange the mannequin’s layers. The primary one, a variational-dense layer, has a single unit. The following distribution layer then takes that unit’s output and makes use of it for the imply of a standard distribution – whereas the dimensions of that Regular is fastened at 1:
You’ll have seen one argument to layer_dense_variational we haven’t mentioned but, kl_weight.
That is used to scale the contribution to the full lack of the KL divergence, and usually ought to equal one over the variety of information factors.
Coaching the mannequin is simple. As customers, we solely specify the damaging log chance a part of the loss; the KL divergence half is taken care of transparently by the framework.
Due to the stochasticity inherent in a variational-dense layer, every time we name this mannequin, we acquire completely different outcomes: completely different regular distributions, on this case.
To acquire the uncertainty estimates we’re on the lookout for, we due to this fact name the mannequin a bunch of instances – 100, say:
yhats <- purrr::map(1:100, operate(x) mannequin(tf$fixed(x_test)))We will now plot these 100 predictions – traces, on this case, as there aren’t any nonlinearities:
means <-
purrr::map(yhats, purrr::compose(as.matrix, tfd_mean)) %>% abind::abind()
traces <- information.body(cbind(x_test, means)) %>%
collect(key = run, worth = worth,-X1)
imply <- apply(means, 1, imply)
ggplot(information.body(x = x, y = y, imply = as.numeric(imply)), aes(x, y)) +
geom_point() +
geom_line(aes(x = x_test, y = imply), shade = "violet", dimension = 1.5) +
geom_line(
information = traces,
aes(x = X1, y = worth, shade = run),
alpha = 0.3,
dimension = 0.5
) +
theme(legend.place = "none")
Determine 4: Epistemic uncertainty on simulated information, utilizing linear activation within the variational-dense layer.
What we see listed here are primarily completely different fashions, in step with the assumptions constructed into the structure. What we’re not accounting for is the unfold within the information. Can we do each? We will; however first let’s touch upon just a few decisions that have been made and see how they have an effect on the outcomes.
To stop this publish from rising to infinite dimension, we’ve kept away from performing a scientific experiment; please take what follows not as generalizable statements, however as tips to issues it would be best to consider in your individual ventures. Particularly, every (hyper-)parameter just isn’t an island; they might work together in unexpected methods.
After these phrases of warning, listed here are some issues we seen.
- One query you would possibly ask: Earlier than, within the aleatoric uncertainty setup, we added a further dense layer to the mannequin, with
reluactivation. What if we did this right here?
Firstly, we’re not including any extra, non-variational layers with a view to maintain the setup “totally Bayesian” – we wish priors at each stage. As to utilizingreluinlayer_dense_variational, we did strive that, and the outcomes look fairly related:

Determine 5: Epistemic uncertainty on simulated information, utilizing relu activation within the variational-dense layer.
Nonetheless, issues look fairly completely different if we drastically scale back coaching time… which brings us to the subsequent remark.
- Not like within the aleatoric setup, the variety of coaching epochs matter rather a lot. If we prepare, quote unquote, too lengthy, the posterior estimates will get nearer and nearer to the posterior imply: we lose uncertainty. What occurs if we prepare “too quick” is much more notable. Listed here are the outcomes for the linear-activation in addition to the relu-activation instances:

Determine 6: Epistemic uncertainty on simulated information if we prepare for 100 epochs solely. Left: linear activation. Proper: relu activation.
Apparently, each mannequin households look very completely different now, and whereas the linear-activation household appears to be like extra cheap at first, it nonetheless considers an general damaging slope in step with the information.
So what number of epochs are “lengthy sufficient”? From remark, we’d say {that a} working heuristic ought to in all probability be based mostly on the speed of loss discount. However definitely, it’ll make sense to strive completely different numbers of epochs and test the impact on mannequin conduct. As an apart, monitoring estimates over coaching time could even yield vital insights into the assumptions constructed right into a mannequin (e.g., the impact of various activation features).
-
As vital because the variety of epochs skilled, and related in impact, is the studying charge. If we change the educational charge on this setup by
0.001, outcomes will look just like what we noticed above for theepochs = 100case. Once more, we’ll need to strive completely different studying charges and ensure we prepare the mannequin “to completion” in some cheap sense. -
To conclude this part, let’s shortly have a look at what occurs if we fluctuate two different parameters. What if the prior have been non-trainable (see the commented line above)? And what if we scaled the significance of the KL divergence (
kl_weightinlayer_dense_variational’s argument record) in another way, changingkl_weight = 1/nbykl_weight = 1(or equivalently, eradicating it)? Listed here are the respective outcomes for an otherwise-default setup. They don’t lend themselves to generalization – on completely different (e.g., larger!) datasets the outcomes will most definitely look completely different – however positively attention-grabbing to look at.

Determine 7: Epistemic uncertainty on simulated information. Left: kl_weight = 1. Proper: prior non-trainable.
Now let’s come again to the query: We’ve modeled unfold within the information, we’ve peeked into the center of the mannequin, – can we do each on the identical time?
We will, if we mix each approaches. We add a further unit to the variational-dense layer and use this to be taught the variance: as soon as for every “sub-model” contained within the mannequin.
Combining each aleatoric and epistemic uncertainty
Reusing the prior and posterior from above, that is how the ultimate mannequin appears to be like:
mannequin <- keras_model_sequential() %>%
layer_dense_variational(
models = 2,
make_posterior_fn = posterior_mean_field,
make_prior_fn = prior_trainable,
kl_weight = 1 / n
) %>%
layer_distribution_lambda(operate(x)
tfd_normal(loc = x[, 1, drop = FALSE],
scale = 1e-3 + tf$math$softplus(0.01 * x[, 2, drop = FALSE])
)
)We prepare this mannequin identical to the epistemic-uncertainty just one. We then acquire a measure of uncertainty per predicted line. Or within the phrases we used above, we now have an ensemble of fashions every with its personal indication of unfold within the information. Here’s a means we might show this – every coloured line is the imply of a distribution, surrounded by a confidence band indicating +/- two customary deviations.
yhats <- purrr::map(1:100, operate(x) mannequin(tf$fixed(x_test)))
means <-
purrr::map(yhats, purrr::compose(as.matrix, tfd_mean)) %>% abind::abind()
sds <-
purrr::map(yhats, purrr::compose(as.matrix, tfd_stddev)) %>% abind::abind()
means_gathered <- information.body(cbind(x_test, means)) %>%
collect(key = run, worth = mean_val,-X1)
sds_gathered <- information.body(cbind(x_test, sds)) %>%
collect(key = run, worth = sd_val,-X1)
traces <-
means_gathered %>% inner_join(sds_gathered, by = c("X1", "run"))
imply <- apply(means, 1, imply)
ggplot(information.body(x = x, y = y, imply = as.numeric(imply)), aes(x, y)) +
geom_point() +
theme(legend.place = "none") +
geom_line(aes(x = x_test, y = imply), shade = "violet", dimension = 1.5) +
geom_line(
information = traces,
aes(x = X1, y = mean_val, shade = run),
alpha = 0.6,
dimension = 0.5
) +
geom_ribbon(
information = traces,
aes(
x = X1,
ymin = mean_val - 2 * sd_val,
ymax = mean_val + 2 * sd_val,
group = run
),
alpha = 0.05,
fill = "gray",
inherit.aes = FALSE
)
Determine 8: Displaying each epistemic and aleatoric uncertainty on the simulated dataset.
Good! This appears to be like like one thing we might report.
As you may think, this mannequin, too, is delicate to how lengthy (assume: variety of epochs) or how briskly (assume: studying charge) we prepare it. And in comparison with the epistemic-uncertainty solely mannequin, there may be a further option to be made right here: the scaling of the earlier layer’s activation – the 0.01 within the scale argument to tfd_normal:
scale = 1e-3 + tf$math$softplus(0.01 * x[, 2, drop = FALSE])Protecting the whole lot else fixed, right here we fluctuate that parameter between 0.01 and 0.05:

Determine 9: Epistemic plus aleatoric uncertainty on the simulated dataset: Various the dimensions argument.
Evidently, that is one other parameter we ought to be ready to experiment with.
Now that we’ve launched all three kinds of presenting uncertainty – aleatoric solely, epistemic solely, or each – let’s see them on the aforementioned Mixed Cycle Energy Plant Information Set. Please see our earlier publish on uncertainty for a fast characterization, in addition to visualization, of the dataset.
Mixed Cycle Energy Plant Information Set
To maintain this publish at a digestible size, we’ll chorus from making an attempt as many alternate options as with the simulated information and primarily stick with what labored properly there. This must also give us an thought of how properly these “defaults” generalize. We individually examine two situations: The one-predictor setup (utilizing every of the 4 obtainable predictors alone), and the entire one (utilizing all 4 predictors without delay).
The dataset is loaded simply as within the earlier publish.
First we have a look at the single-predictor case, ranging from aleatoric uncertainty.
Single predictor: Aleatoric uncertainty
Right here is the “default” aleatoric mannequin once more. We additionally duplicate the plotting code right here for the reader’s comfort.
n <- nrow(X_train) # 7654
n_epochs <- 10 # we'd like fewer epochs as a result of the dataset is a lot larger
batch_size <- 100
learning_rate <- 0.01
# variable to suit - change to 2,3,4 to get the opposite predictors
i <- 1
mannequin <- keras_model_sequential() %>%
layer_dense(models = 16, activation = "relu") %>%
layer_dense(models = 2, activation = "linear") %>%
layer_distribution_lambda(operate(x)
tfd_normal(loc = x[, 1, drop = FALSE],
scale = tf$math$softplus(x[, 2, drop = FALSE])
)
)
negloglik <- operate(y, mannequin) - (mannequin %>% tfd_log_prob(y))
mannequin %>% compile(optimizer = optimizer_adam(lr = learning_rate), loss = negloglik)
hist <-
mannequin %>% match(
X_train[, i, drop = FALSE],
y_train,
validation_data = record(X_val[, i, drop = FALSE], y_val),
epochs = n_epochs,
batch_size = batch_size
)
yhat <- mannequin(tf$fixed(X_val[, i, drop = FALSE]))
imply <- yhat %>% tfd_mean()
sd <- yhat %>% tfd_stddev()
ggplot(information.body(
x = X_val[, i],
y = y_val,
imply = as.numeric(imply),
sd = as.numeric(sd)
),
aes(x, y)) +
geom_point() +
geom_line(aes(x = x, y = imply), shade = "violet", dimension = 1.5) +
geom_ribbon(aes(
x = x,
ymin = imply - 2 * sd,
ymax = imply + 2 * sd
),
alpha = 0.4,
fill = "gray")How properly does this work?

Determine 10: Aleatoric uncertainty on the Mixed Cycle Energy Plant Information Set; single predictors.
This appears to be like fairly good we’d say! How about epistemic uncertainty?
Single predictor: Epistemic uncertainty
Right here’s the code:
posterior_mean_field <-
operate(kernel_size,
bias_size = 0,
dtype = NULL) {
n <- kernel_size + bias_size
c <- log(expm1(1))
keras_model_sequential(record(
layer_variable(form = 2 * n, dtype = dtype),
layer_distribution_lambda(
make_distribution_fn = operate(t) {
tfd_independent(tfd_normal(
loc = t[1:n],
scale = 1e-5 + tf$nn$softplus(c + t[(n + 1):(2 * n)])
), reinterpreted_batch_ndims = 1)
}
)
))
}
prior_trainable <-
operate(kernel_size,
bias_size = 0,
dtype = NULL) {
n <- kernel_size + bias_size
keras_model_sequential() %>%
layer_variable(n, dtype = dtype, trainable = TRUE) %>%
layer_distribution_lambda(operate(t) {
tfd_independent(tfd_normal(loc = t, scale = 1),
reinterpreted_batch_ndims = 1)
})
}
mannequin <- keras_model_sequential() %>%
layer_dense_variational(
models = 1,
make_posterior_fn = posterior_mean_field,
make_prior_fn = prior_trainable,
kl_weight = 1 / n,
activation = "linear",
) %>%
layer_distribution_lambda(operate(x)
tfd_normal(loc = x, scale = 1))
negloglik <- operate(y, mannequin) - (mannequin %>% tfd_log_prob(y))
mannequin %>% compile(optimizer = optimizer_adam(lr = learning_rate), loss = negloglik)
hist <-
mannequin %>% match(
X_train[, i, drop = FALSE],
y_train,
validation_data = record(X_val[, i, drop = FALSE], y_val),
epochs = n_epochs,
batch_size = batch_size
)
yhats <- purrr::map(1:100, operate(x)
yhat <- mannequin(tf$fixed(X_val[, i, drop = FALSE])))
means <-
purrr::map(yhats, purrr::compose(as.matrix, tfd_mean)) %>% abind::abind()
traces <- information.body(cbind(X_val[, i], means)) %>%
collect(key = run, worth = worth,-X1)
imply <- apply(means, 1, imply)
ggplot(information.body(x = X_val[, i], y = y_val, imply = as.numeric(imply)), aes(x, y)) +
geom_point() +
geom_line(aes(x = X_val[, i], y = imply), shade = "violet", dimension = 1.5) +
geom_line(
information = traces,
aes(x = X1, y = worth, shade = run),
alpha = 0.3,
dimension = 0.5
) +
theme(legend.place = "none")And that is the consequence.

Determine 11: Epistemic uncertainty on the Mixed Cycle Energy Plant Information Set; single predictors.
As with the simulated information, the linear fashions appears to “do the proper factor”. And right here too, we expect we’ll need to increase this with the unfold within the information: Thus, on to means three.
Single predictor: Combining each sorts
Right here we go. Once more, posterior_mean_field and prior_trainable look identical to within the epistemic-only case.
mannequin <- keras_model_sequential() %>%
layer_dense_variational(
models = 2,
make_posterior_fn = posterior_mean_field,
make_prior_fn = prior_trainable,
kl_weight = 1 / n,
activation = "linear"
) %>%
layer_distribution_lambda(operate(x)
tfd_normal(loc = x[, 1, drop = FALSE],
scale = 1e-3 + tf$math$softplus(0.01 * x[, 2, drop = FALSE])))
negloglik <- operate(y, mannequin)
- (mannequin %>% tfd_log_prob(y))
mannequin %>% compile(optimizer = optimizer_adam(lr = learning_rate), loss = negloglik)
hist <-
mannequin %>% match(
X_train[, i, drop = FALSE],
y_train,
validation_data = record(X_val[, i, drop = FALSE], y_val),
epochs = n_epochs,
batch_size = batch_size
)
yhats <- purrr::map(1:100, operate(x)
mannequin(tf$fixed(X_val[, i, drop = FALSE])))
means <-
purrr::map(yhats, purrr::compose(as.matrix, tfd_mean)) %>% abind::abind()
sds <-
purrr::map(yhats, purrr::compose(as.matrix, tfd_stddev)) %>% abind::abind()
means_gathered <- information.body(cbind(X_val[, i], means)) %>%
collect(key = run, worth = mean_val,-X1)
sds_gathered <- information.body(cbind(X_val[, i], sds)) %>%
collect(key = run, worth = sd_val,-X1)
traces <-
means_gathered %>% inner_join(sds_gathered, by = c("X1", "run"))
imply <- apply(means, 1, imply)
#traces <- traces %>% filter(run=="X3" | run =="X4")
ggplot(information.body(x = X_val[, i], y = y_val, imply = as.numeric(imply)), aes(x, y)) +
geom_point() +
theme(legend.place = "none") +
geom_line(aes(x = X_val[, i], y = imply), shade = "violet", dimension = 1.5) +
geom_line(
information = traces,
aes(x = X1, y = mean_val, shade = run),
alpha = 0.2,
dimension = 0.5
) +
geom_ribbon(
information = traces,
aes(
x = X1,
ymin = mean_val - 2 * sd_val,
ymax = mean_val + 2 * sd_val,
group = run
),
alpha = 0.01,
fill = "gray",
inherit.aes = FALSE
)And the output?

Determine 12: Mixed uncertainty on the Mixed Cycle Energy Plant Information Set; single predictors.
This appears to be like helpful! Let’s wrap up with our ultimate take a look at case: Utilizing all 4 predictors collectively.
All predictors
The coaching code used on this situation appears to be like identical to earlier than, aside from our feeding all predictors to the mannequin. For plotting, we resort to displaying the primary principal element on the x-axis – this makes the plots look noisier than earlier than. We additionally show fewer traces for the epistemic and epistemic-plus-aleatoric instances (20 as a substitute of 100). Listed here are the outcomes:

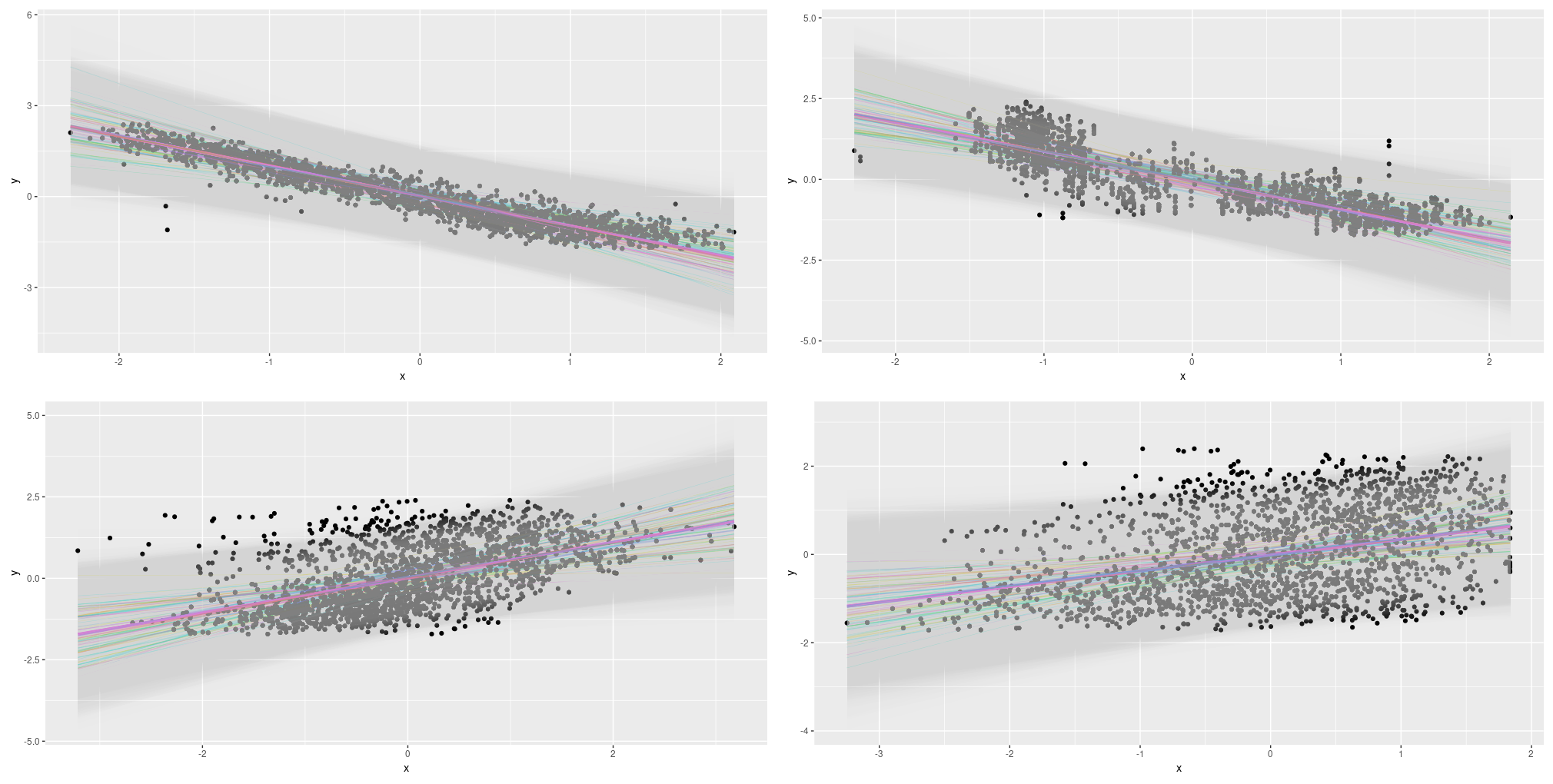
Determine 13: Uncertainty (aleatoric, epistemic, each) on the Mixed Cycle Energy Plant Information Set; all predictors.
Conclusion
The place does this depart us? In comparison with the learnable-dropout method described within the prior publish, the way in which introduced here’s a lot simpler, quicker, and extra intuitively comprehensible.
The strategies per se are that straightforward to make use of that on this first introductory publish, we might afford to discover alternate options already: one thing we had no time to do in that earlier exposition.
In actual fact, we hope this publish leaves you ready to do your individual experiments, by yourself information.
Clearly, you’ll have to make choices, however isn’t that the way in which it’s in information science? There’s no means round making choices; we simply ought to be ready to justify them …
Thanks for studying!